Media Manager
Media Files
Files in training

- 01_introductiontor.html.zip
- 2022/04/21 10:49
- 518 KB

- 01_introductiontor_cor.html.zip
- 2022/04/29 10:04
- 521.4 KB

- 02_graphicswithggplot2.html.zip
- 2022/04/29 10:04
- 1.5 MB

- 02_graphicswithggplot2_cor.html.zip
- 2022/05/06 09:52
- 2.3 MB

- 03_generatereportwithrmarkdown.html.zip
- 2022/05/06 09:50
- 373.1 KB

- 03_generatereportwithrmarkdown_cor.html.zip
- 2022/05/06 15:12
- 373.9 KB

- 1701ngsbioinfo_0_intro.pptx.pdf
- 2019/07/22 11:07
- 41 KB

- 1701ngsbioinfo_02a_galaxy.pdf
- 2019/07/22 11:07
- 5.8 MB

- 1701ngsbioinfo_02b_galaxy_correction.pdf
- 2019/07/22 11:07
- 334 KB

- 1701ngsbioinfo_08a_biomart.pdf
- 2019/07/22 11:07
- 4.3 MB

- 1701ngsbioinfo_08b_biomart_correction.pdf
- 2019/07/22 11:07
- 368.2 KB

- 1701ngsbioinfo_1_ngs.pptx.pdf
- 2019/07/22 11:07
- 19.6 MB

- 1701ngsbioinfo_3_rnaseq_preplib_expdesign.pptx.pdf
- 2019/07/22 11:07
- 7.9 MB

- 1701ngsbioinfo_4_intro_practical.pptx.pdf
- 2019/07/22 11:07
- 89.9 KB

- 1701ngsbioinfo_5_qc.pptx.pdf
- 2019/07/22 11:07
- 4.2 MB

- 1701ngsbioinfo_6_mapping.pptx.pdf
- 2019/07/22 11:07
- 5.6 MB

- 1701ngsbioinfo_6_mapping_corrections.pptx.pdf
- 2019/07/22 11:07
- 4.1 MB

- 1701ngsbioinfo_7_rnaseq_analysis.pptx.pdf
- 2019/07/22 11:07
- 5 MB

- 1701ngsbioinfo_7_rnaseq_analysis_corrections.pptx.pdf
- 2019/07/22 11:07
- 907 KB

- 1701ngsbioinfo_9_functional_analysis.pptx.pdf
- 2019/07/22 11:07
- 1 MB

- 1701ngsbioinfo_9_functional_analysis_corrections.pptx.pdf
- 2019/07/22 11:07
- 1.4 MB

- 1701ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2019/07/22 11:07
- 2.1 MB

- 1701ngsbioinfo_11a_chipseq_mapping_visualization.pdf
- 2019/07/22 11:07
- 310.9 KB

- 1701ngsbioinfo_11b_chipseq_mapping_visualization_correction.pdf
- 2019/07/22 11:07
- 153.3 KB

- 1701ngsbioinfo_12a_chipseq_peakcalling.pdf
- 2019/07/22 11:07
- 1.5 MB

- 1701ngsbioinfo_12b_chipseq_peakcalling_correction.pdf
- 2019/07/22 11:07
- 594.6 KB

- 1701ngsbioinfo_13a_chipseq_peakanalysis.pdf
- 2019/07/22 11:07
- 2.4 MB

- 1701ngsbioinfo_13b_chipseq_peakanalysis_correction.pdf
- 2019/07/22 11:07
- 256.3 KB

- 1701ngsbioinfo_14a_workflows.pdf
- 2019/07/22 11:07
- 2.4 MB

- 1701ngsbioinfo_14b_workflows_correction.pdf
- 2019/07/22 11:07
- 1.2 MB

- 1701ngsbioinfo_15a_chipseq_rnaseq_correlation.pdf
- 2019/07/22 11:07
- 45.3 KB

- 1701ngsbioinfo_15b_chipseq_rnaseq_correlation_correction.pdf
- 2019/07/22 11:07
- 395.2 KB

- 1709ngsbioinfo_02a_galaxy.pdf
- 2019/07/22 11:07
- 6.9 MB

- 1709ngsbioinfo_02b_galaxy_correction.pdf
- 2019/07/22 11:07
- 731.9 KB

- 1709ngsbioinfo_08a_biomart.pdf
- 2019/07/22 11:07
- 4.8 MB

- 1709ngsbioinfo_08b_biomart_correction.pdf
- 2019/07/22 11:07
- 1 MB

- 1709ngsbioinfo_1_ngs.pdf
- 2020/03/30 14:39
- 20.7 MB

- 1709ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2019/07/22 11:07
- 7.9 MB

- 1709ngsbioinfo_4_intro_practical.pdf
- 2019/07/22 11:07
- 39.2 KB

- 1709ngsbioinfo_5_qc.pdf
- 2019/07/22 11:07
- 4.4 MB

- 1709ngsbioinfo_6_mapping.pdf
- 2019/07/22 11:07
- 5.8 MB

- 1709ngsbioinfo_6_mapping_corrections.pdf
- 2019/07/22 11:07
- 3.7 MB

- 1709ngsbioinfo_7_rnaseq_analysis.pdf
- 2019/07/22 11:07
- 5.3 MB

- 1709ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2019/07/22 11:07
- 930.5 KB

- 1709ngsbioinfo_9_functional_analysis.pdf
- 2019/07/22 11:07
- 1.2 MB

- 1709ngsbioinfo_9_functional_analysis_corrections.pdf
- 2019/07/22 11:07
- 1.8 MB

- 1709ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2019/07/22 11:07
- 3 MB

- 1709ngsbioinfo_11a_chipseq_mapping_visualization.pdf
- 2019/07/22 11:07
- 554.7 KB

- 1709ngsbioinfo_11b_chipseq_mapping_visualization_correction.pdf
- 2019/07/22 11:07
- 480.5 KB

- 1709ngsbioinfo_12a_chipseq_peakcalling.pdf
- 2019/07/22 11:07
- 2.2 MB

- 1709ngsbioinfo_12b_chipseq_peakcalling_correction.pdf
- 2019/07/22 11:07
- 1.1 MB

- 1709ngsbioinfo_13a_chipseq_peakanalysis.pdf
- 2019/07/22 11:07
- 3.6 MB

- 1709ngsbioinfo_13b_chipseq_peakanalysis_correction.pdf
- 2019/07/22 11:07
- 706.3 KB

- 1709ngsbioinfo_14a_workflows.pdf
- 2019/07/22 11:07
- 3 MB

- 1709ngsbioinfo_14b_workflows_correction.pdf
- 2019/07/22 11:07
- 2.6 MB

- 1709ngsbioinfo_15a_chipseq_rnaseq_correlation.pdf
- 2019/07/22 11:07
- 110.3 KB

- 1709ngsbioinfo_15b_chipseq_rnaseq_correlation_correction.pdf
- 2019/07/22 11:07
- 588 KB

- 1903ngsbioinfo_02a_galaxy.pdf
- 2019/07/22 11:07
- 2.4 MB

- 1903ngsbioinfo_02b_galaxy_correction.pdf
- 2019/07/22 11:07
- 358.1 KB

- 1903ngsbioinfo_08a_biomart.pdf
- 2019/07/22 11:07
- 956.7 KB

- 1903ngsbioinfo_08b_biomart_correction.pdf
- 2019/07/22 11:07
- 169.1 KB

- 1903ngsbioinfo_1_ngs.pdf
- 2019/07/22 11:07
- 23.4 MB

- 1903ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2019/07/22 11:07
- 10.8 MB

- 1903ngsbioinfo_4_intro_practical.pdf
- 2019/07/22 11:07
- 100.8 KB

- 1903ngsbioinfo_5_qc.pdf
- 2019/07/22 11:07
- 13.9 MB

- 1903ngsbioinfo_6_mapping.pdf
- 2019/07/22 11:07
- 12.6 MB

- 1903ngsbioinfo_6_mapping_corrections.pdf
- 2019/07/22 11:07
- 11.1 MB

- 1903ngsbioinfo_7_rnaseq_analysis.pdf
- 2019/07/22 11:07
- 23.7 MB

- 1903ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2019/07/22 11:07
- 4.2 MB

- 1903ngsbioinfo_9_functional_analysis.pdf
- 2019/07/22 11:07
- 4.8 MB

- 1903ngsbioinfo_9_functional_analysis_corrections.pdf
- 2019/07/22 11:07
- 7.8 MB

- 1903ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2019/07/22 11:07
- 843.5 KB

- 1903ngsbioinfo_11a_chipseq_mapping_visualization.pdf
- 2019/07/22 11:07
- 202.6 KB

- 1903ngsbioinfo_11b_chipseq_mapping_visualization_correction.pdf
- 2019/07/22 11:07
- 68.8 KB

- 1903ngsbioinfo_12a_chipseq_peakcalling.pdf
- 2019/07/22 11:07
- 2.1 MB

- 1903ngsbioinfo_12b_chipseq_peakcalling_correction.pdf
- 2019/07/22 11:07
- 92.8 KB

- 1903ngsbioinfo_13a_chipseq_peakanalysis.pdf
- 2019/07/22 11:07
- 1.6 MB

- 1903ngsbioinfo_13b_chipseq_peakanalysis_correction.pdf
- 2019/07/22 11:07
- 386.8 KB

- 1903ngsbioinfo_14a_workflows.pdf
- 2019/07/22 11:07
- 1.5 MB

- 1903ngsbioinfo_14b_workflows_correction.pdf
- 2019/07/22 11:07
- 556.7 KB

- 1903ngsbioinfo_15a_chipseq_rnaseq_correlation.pdf
- 2019/07/22 11:07
- 37.6 KB

- 1903ngsbioinfo_15b_chipseq_rnaseq_correlation_correction.pdf
- 2019/07/22 11:07
- 229.5 KB

- 1911ngsbioinfo_02a_galaxy.pdf
- 2019/11/19 13:47
- 1.8 MB

- 1911ngsbioinfo_02b_galaxy_correction.pdf
- 2019/11/19 13:47
- 227.4 KB

- 1911ngsbioinfo_08a_biomart.pdf
- 2019/11/19 13:47
- 1.1 MB

- 1911ngsbioinfo_08b_biomart_correction.pdf
- 2019/11/19 13:47
- 167.7 KB

- 1911ngsbioinfo_1_ngs.pdf
- 2019/11/19 13:47
- 16.5 MB

- 1911ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2019/11/19 13:47
- 7.6 MB

- 1911ngsbioinfo_4_intro_practical.pdf
- 2019/11/19 13:47
- 98.2 KB

- 1911ngsbioinfo_5_qc.pdf
- 2019/11/19 13:47
- 13.6 MB

- 1911ngsbioinfo_6_mapping.pdf
- 2019/11/19 13:47
- 13.4 MB

- 1911ngsbioinfo_6_mapping_corrections.pdf
- 2019/11/19 13:47
- 12 MB

- 1911ngsbioinfo_7_rnaseq_analysis.pdf
- 2019/11/19 13:47
- 24 MB

- 1911ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2019/11/19 13:47
- 4.2 MB

- 1911ngsbioinfo_9_functional_analysis.pdf
- 2019/11/19 13:47
- 4.8 MB

- 1911ngsbioinfo_9_functional_analysis_corrections.pdf
- 2019/11/19 13:47
- 7.8 MB

- 1911ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2019/11/19 13:47
- 677.7 KB

- 1911ngsbioinfo_11a_chipseq_dataanalysis.pdf
- 2019/11/19 13:47
- 11.9 MB

- 1911ngsbioinfo_11b_chipseq_dataanalysis_correction.pdf
- 2019/11/19 13:47
- 474.1 KB

- 1911ngsbioinfo_12_workflows.pdf
- 2019/11/19 13:47
- 2.1 MB

- 1911ngsbioinfo_13a_chipseq_rnaseq_correlation.pdf
- 2019/11/19 13:47
- 37.4 KB

- 1911ngsbioinfo_13b_chipseq_rnaseq_correlation_correction.pdf
- 2019/11/19 13:47
- 237.3 KB

- 2009ngsbioinfo_02a_galaxy.pdf
- 2020/09/15 21:07
- 29.6 MB

- 2009ngsbioinfo_02b_galaxy_correction.pdf
- 2020/09/15 21:07
- 2.1 MB

- 2009ngsbioinfo_08a_biomart.pdf
- 2020/09/15 21:07
- 11 MB

- 2009ngsbioinfo_08b_biomart_correction.pdf
- 2020/09/15 21:07
- 1.1 MB

- 2009ngsbioinfo_1_ngs.pdf
- 2020/09/08 16:39
- 15.9 MB

- 2009ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2020/09/08 16:39
- 7.5 MB

- 2009ngsbioinfo_4_intro_practical.pdf
- 2020/09/08 16:39
- 98.1 KB

- 2009ngsbioinfo_5_qc.pdf
- 2020/09/08 16:39
- 13.2 MB

- 2009ngsbioinfo_6_mapping.pdf
- 2020/09/08 16:39
- 14 MB

- 2009ngsbioinfo_6_mapping_corrections.pdf
- 2020/09/08 16:39
- 13.2 MB

- 2009ngsbioinfo_7_rnaseq_analysis.pdf
- 2020/09/08 16:39
- 27.7 MB

- 2009ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2020/09/08 16:39
- 4.2 MB

- 2009ngsbioinfo_9_functional_analysis.pdf
- 2020/09/08 16:39
- 6.5 MB

- 2009ngsbioinfo_9_functional_analysis_corrections.pdf
- 2020/09/08 16:39
- 10.1 MB

- 2009ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2020/09/15 21:07
- 6.2 MB

- 2009ngsbioinfo_11a_chipseq_dataanalysis.pdf
- 2020/09/15 21:08
- 34.8 MB

- 2009ngsbioinfo_11b_chipseq_dataanalysis_correction.pdf
- 2020/09/15 21:07
- 8.7 MB

- 2009ngsbioinfo_12_workflows.pdf
- 2020/09/15 21:08
- 26.3 MB

- 2009ngsbioinfo_13a_chipseq_rnaseq_correlation.pdf
- 2020/09/15 21:07
- 169.3 KB

- 2009ngsbioinfo_13b_chipseq_rnaseq_correlation_correction.pdf
- 2020/09/15 21:07
- 2.4 MB

- 2016phdprogramme_chipseq.pdf
- 2019/07/22 11:07
- 9.4 MB

- 2106ngsbioinfo_02a_galaxy.pdf
- 2021/06/16 21:43
- 8 MB

- 2106ngsbioinfo_02b_galaxy_correction.pdf
- 2021/06/16 21:43
- 907.8 KB

- 2106ngsbioinfo_08a_biomart.pdf
- 2021/06/16 21:43
- 6.4 MB

- 2106ngsbioinfo_08b_biomart_correction.pdf
- 2021/06/16 21:43
- 241.8 KB

- 2106ngsbioinfo_1_ngs.pdf
- 2021/06/07 12:16
- 5.2 MB

- 2106ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2021/06/07 12:18
- 10.5 MB

- 2106ngsbioinfo_4_intro_practical.pdf
- 2021/06/07 12:19
- 84.3 KB

- 2106ngsbioinfo_5_qc.pdf
- 2021/06/07 12:20
- 5.1 MB

- 2106ngsbioinfo_6_mapping.pdf
- 2021/06/07 12:20
- 5.7 MB

- 2106ngsbioinfo_6_mapping_corrections.pdf
- 2021/06/07 12:20
- 5.1 MB

- 2106ngsbioinfo_7_rnaseq_analysis.pdf
- 2021/06/07 12:22
- 8.9 MB

- 2106ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2021/06/07 12:22
- 2 MB

- 2106ngsbioinfo_9_functional_analysis.pdf
- 2021/06/07 12:22
- 2.5 MB

- 2106ngsbioinfo_9_functional_analysis_corrections.pdf
- 2021/06/07 12:22
- 2.6 MB

- 2106ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2021/06/16 21:43
- 6.3 MB

- 2106ngsbioinfo_11a_chipseq_dataanalysis.pdf
- 2021/06/16 21:43
- 26.1 MB

- 2106ngsbioinfo_11b_chipseq_dataanalysis_correction.pdf
- 2021/06/16 21:43
- 3.3 MB

- 2106ngsbioinfo_12_workflows.pdf
- 2021/06/16 21:43
- 13.1 MB

- 2106ngsbioinfo_13a_chipseq_rnaseq_correlation.pdf
- 2021/06/16 21:43
- 147.2 KB

- 2106ngsbioinfo_13b_chipseq_rnaseq_correlation_correction.pdf
- 2021/06/16 21:43
- 1.4 MB

- 2203ngsbioinfo_0_intro.pdf
- 2022/02/24 14:54
- 212.7 KB

- 2203ngsbioinfo_02a_galaxy.pdf
- 2022/03/01 21:14
- 12.4 MB

- 2203ngsbioinfo_02b_galaxy_correction.pdf
- 2022/03/01 21:14
- 1.4 MB

- 2203ngsbioinfo_08a_biomart.pdf
- 2022/03/01 21:14
- 5.8 MB

- 2203ngsbioinfo_08b_biomart_correction.pdf
- 2022/03/01 21:14
- 4.3 MB

- 2203ngsbioinfo_1_ngs.pdf
- 2022/03/10 17:08
- 4.7 MB

- 2203ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2022/03/10 17:08
- 6.8 MB

- 2203ngsbioinfo_4_intro_practical.pdf
- 2022/03/10 17:08
- 83.5 KB

- 2203ngsbioinfo_5_qc.pdf
- 2022/03/10 17:08
- 4.4 MB

- 2203ngsbioinfo_6_mapping.pdf
- 2022/03/10 17:08
- 8.2 MB

- 2203ngsbioinfo_6_mapping_corrections.pdf
- 2022/03/10 17:08
- 6.1 MB

- 2203ngsbioinfo_7_rnaseq_analysis.pdf
- 2022/03/10 17:08
- 12.9 MB

- 2203ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2022/03/10 17:08
- 1.6 MB

- 2203ngsbioinfo_9_functional_analysis.pdf
- 2022/03/10 17:08
- 2.8 MB

- 2203ngsbioinfo_9_functional_analysis_corrections.pdf
- 2022/03/10 17:08
- 3.3 MB

- 2203ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2022/03/24 11:25
- 7.5 MB

- 2203ngsbioinfo_10_chipseq_libprep_expdesign.pptx
- 2022/03/14 09:31
- 1.9 MB

- 2203ngsbioinfo_11a_chipseq_dataanalysis.pdf
- 2022/02/08 17:09
- 11.4 MB

- 2203ngsbioinfo_11a_chipseq_dataanalysis_compressed.pdf
- 2022/03/01 21:14
- 4.4 MB

- 2203ngsbioinfo_11b_chipseq_dataanalysis_correction.pdf
- 2022/03/01 21:14
- 5.3 MB

- 2203ngsbioinfo_12_workflows.pdf
- 2022/02/08 17:09
- 22.8 MB

- 2203ngsbioinfo_12_workflows_compressed.pdf
- 2022/03/01 21:14
- 2.6 MB

- 2203ngsbioinfo_13a_chipseq_rnaseq_correlation.pdf
- 2022/03/01 21:15
- 3.4 MB

- 2203ngsbioinfo_13b_chipseq_rnaseq_correlation_correction.pdf
- 2022/03/01 21:15
- 638.1 KB

- 2305ngsbioinfo_02a_galaxy.pdf
- 2023/04/21 10:12
- 6.9 MB

- 2305ngsbioinfo_02b_galaxy_correction.pdf
- 2023/04/21 10:12
- 1.4 MB

- 2305ngsbioinfo_08a_biomart.pdf
- 2023/04/21 10:12
- 4.9 MB

- 2305ngsbioinfo_08b_biomart_correction.pdf
- 2023/04/21 10:12
- 6 MB

- 2305ngsbioinfo_1_ngs.pdf
- 2023/05/22 10:52
- 2.6 MB

- 2305ngsbioinfo_3_rnaseq_preplib_expdesign.pdf
- 2023/05/22 10:52
- 2.5 MB

- 2305ngsbioinfo_4_intro_practical.pdf
- 2023/05/22 10:52
- 83.3 KB

- 2305ngsbioinfo_5_qc.pdf
- 2023/05/22 10:52
- 2.7 MB

- 2305ngsbioinfo_6_mapping.pdf
- 2023/05/22 10:52
- 5.5 MB

- 2305ngsbioinfo_6_mapping_corrections.pdf
- 2023/05/16 16:05
- 3.5 MB

- 2305ngsbioinfo_7_rnaseq_analysis.pdf
- 2023/05/22 10:52
- 7.7 MB

- 2305ngsbioinfo_7_rnaseq_analysis_corrections.pdf
- 2023/05/22 10:52
- 1 MB

- 2305ngsbioinfo_9_functional_analysis.pdf
- 2023/05/22 10:52
- 1.6 MB

- 2305ngsbioinfo_9_functional_analysis_corrections.pdf
- 2023/05/22 10:52
- 1.2 MB

- 2305ngsbioinfo_10_chipseq_libprep_expdesign.pdf
- 2023/04/21 10:12
- 4.6 MB

- 2305ngsbioinfo_11a_chipseq_dataanalysis.pdf
- 2023/04/21 10:12
- 19.9 MB

- 2305ngsbioinfo_11b_chipseq_dataanalysis_correction.pdf
- 2023/04/21 10:12
- 4.4 MB

- 2305ngsbioinfo_12_workflows.pdf
- 2023/04/21 10:12
- 23.4 MB

- 2305ngsbioinfo_13a_chipseq_rnaseq_correlation.pdf
- 2023/04/21 10:12
- 2.1 MB

- 2305ngsbioinfo_13b_chipseq_rnaseq_correlation_correction.pdf
- 2023/04/21 10:12
- 667.5 KB

- 180116_rnaseq_phdpgr.pdf
- 2019/07/22 11:07
- 10 MB

- 180116_smallrnaseq_phdpgr.pdf
- 2019/07/22 11:07
- 3 MB

- 180323_r_phdpgr.pdf
- 2019/07/22 11:07
- 4 MB

- 180323_r_phdpgr_corrections.pdf
- 2019/07/22 11:07
- 878.2 KB

- 190115_rnaseq_phdpgr.pdf
- 2019/07/22 11:07
- 20 MB

- 190129_ensembl_phdpgr.pdf
- 2019/07/22 11:07
- 34.8 MB

- 190129_ensembl_phdpgr_corrections.pdf
- 2019/07/22 11:07
- 13.8 MB
- bbs5-205codingexons.png
- 1968×1318
- 2022/01/25 10:42
- 419 KB
- bbs5-205exons.png
- 1972×1320
- 2022/01/25 10:42
- 417.9 KB
- bbs5_003_codingexons.png
- 1267×776
- 2019/07/22 11:07
- 252.7 KB
- bbs5_003_exons.png
- 1267×776
- 2019/07/22 11:07
- 249.1 KB
- bbs5_transcripts.png
- 1982×1474
- 2022/01/25 10:42
- 517.7 KB
- brca1_displayvariation.png
- 1968×1488
- 2022/01/25 10:42
- 536.5 KB
- brca1_displayvariationtable1.png
- 269×540
- 2019/07/22 11:07
- 57.2 KB

- brca1_displayvariationtable2.png
- 1194×792
- 2019/07/22 11:07
- 372.6 KB

- brca1_gograph.png
- 1196×793
- 2019/07/22 11:07
- 277.9 KB

- brca1_goterm.png
- 1194×791
- 2019/07/22 11:07
- 433.7 KB

- brca1_rnaseq.png
- 1598×1150
- 2022/01/25 10:51
- 316.2 KB
- brca1_rnaseq1.png
- 1968×1436
- 2022/01/25 10:51
- 499.1 KB
- brca1_rnaseq2.png
- 1992×1488
- 2022/01/25 10:51
- 548.1 KB
- brca1_sequence.png
- 1972×1488
- 2022/01/25 10:55
- 445.1 KB
- brca1_variation.png
- 2290×1512
- 2021/03/09 18:58
- 598.7 KB
- brca1_variationtable.png
- 1193×787
- 2019/07/22 11:07
- 345 KB

- browseucsc.png
- 1039×255
- 2019/07/22 11:07
- 70.4 KB
- changedisplaymode1.png
- 940×691
- 2019/07/22 11:07
- 386.5 KB

- changedisplaymode2.png
- 1044×475
- 2019/07/22 11:07
- 164 KB
- changehistoryname.png
- 1042×766
- 2019/07/22 11:07
- 239.1 KB

- chipseq_ibmp_12juin2015.pdf
- 2019/07/22 11:07
- 10.7 MB

- datafiles.zip
- 2019/07/22 11:07
- 6.4 KB
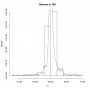
- distance2tss.png
- 434×429
- 2019/07/22 11:07
- 36.2 KB

- ensembl_biomart_2016.pdf
- 2019/07/22 11:07
- 21.5 MB

- ensembl_biomart_2019-compressed.pdf
- 2019/07/22 11:07
- 5.2 MB

- ensembl_biomart_2022_compressed.pdf
- 2022/01/25 11:22
- 3.2 MB

- ensembl_biomart_french.pdf
- 2019/07/22 11:07
- 22.1 MB

- ensembl_biomart_tp_tools.pdf
- 2019/07/22 11:07
- 4 MB
- ensemblcolorcode.png
- 1528×718
- 2022/01/25 10:42
- 178.4 KB

- ensembltranscriptids.txt.zip
- 2019/07/22 11:07
- 2.3 KB

- enstranscriptids.txt.zip
- 2019/07/22 11:07
- 2.8 KB
- humangenomeassembly.png
- 824×722
- 2022/01/25 10:25
- 147.5 KB
- includedeleteddataset.png
- 1043×768
- 2019/07/22 11:07
- 261.5 KB

- introduction_to_chip-seq_data_analysis.pdf
- 2019/07/22 11:07
- 13.7 MB
- managecustomtrack.png
- 1154×383
- 2019/07/22 11:07
- 128.2 KB

- memeresults.png
- 720×378
- 2019/07/22 11:07
- 157.2 KB
- nakedmolerat.png
- 840×486
- 2022/01/25 10:25
- 118.2 KB
- nakedmolerat_genes.png
- 823×634
- 2022/01/25 10:25
- 176.7 KB

- ngsbioinfo_3_intro_practical.pdf
- 2019/07/22 11:07
- 163.7 KB

- ngsbioinfo_4_rnaseq_preplib_expdesign.pdf
- 2019/07/22 11:07
- 7.9 MB

- ngsbioinfo_5_qc.pdf
- 2019/07/22 11:07
- 2.8 MB

- ngsbioinfo_6_mapping.pdf
- 2019/07/22 11:07
- 2.4 MB

- ngsbioinfo_6_mapping_answers.pdf
- 2019/07/22 11:07
- 2.8 MB

- ngsbioinfo_7_rnaseq_analysis.pdf
- 2019/07/22 11:07
- 2.2 MB

- ngsbioinfo_7_rnaseq_analysis_answers.pdf
- 2019/07/22 11:07
- 1.3 MB

- ngsbioinfo_9_chipseq.pdf
- 2019/07/22 11:07
- 15.3 MB

- ngsbioinfo_10_correlation.pdf
- 2019/07/22 11:07
- 7 MB
- numberofgenehuman.png
- 823×697
- 2022/01/25 10:26
- 189.3 KB
- numberofgenehumangrch37.png
- 821×732
- 2022/01/25 10:26
- 176.3 KB
- peaksummit.png
- 1162×762
- 2019/07/22 11:07
- 399.7 KB

- phd_program2019_smallrna.pdf
- 2019/07/22 11:07
- 2.9 MB

- phdprogramme_ensembl_2016.pdf
- 2019/07/22 11:07
- 17.7 MB

- phdprogramme_ensembl_2021.pdf
- 2021/04/08 11:59
- 27 MB
- question5.png
- 1972×1488
- 2022/01/25 10:53
- 441.9 KB

- registration_form_ngsbioinfo19.docx
- 2019/07/22 11:07
- 123.1 KB
- rerunmacs.png
- 400×300
- 2019/07/22 11:07
- 48.1 KB

- resultmacs.png
- 742×402
- 2019/07/22 11:07
- 315.6 KB
- resultmacs2.png
- 629×475
- 2019/07/22 11:07
- 110.8 KB

- resultmacs3.png
- 720×391
- 2019/07/22 11:07
- 161.4 KB

- resultmacs4.png
- 721×417
- 2019/07/22 11:07
- 230.1 KB
- resultrerunmacs.png
- 756×397
- 2019/07/22 11:07
- 313.2 KB

- simitfvssiluc-up.tsv.gz
- 2019/07/22 11:07
- 222.5 KB

- simitfvssiluc-up.tsv.zip
- 2019/07/22 11:07
- 223.1 KB

- simitfvssiluc.up.txt.zip
- 2022/01/25 11:00
- 215.8 KB
- sizechr4.png
- 968×746
- 2022/01/25 10:26
- 159.7 KB

- t.yeravensetal2014.pdf
- 2021/08/02 15:42
- 402.5 KB
- undeletedatasets.png
- 1044×770
- 2019/07/22 11:07
- 244.7 KB
- uploadpeaksummits.png
- 233×330
- 2019/07/22 11:07
- 28.8 KB
File
- Date:
- 2023/05/22 10:52
- Filename:
- 2305ngsbioinfo_6_mapping.pdf
- Size:
- 5MB

